13/02/2026
V 1.0.0 – IUCN Red List eDNA Dashboard Initial Release
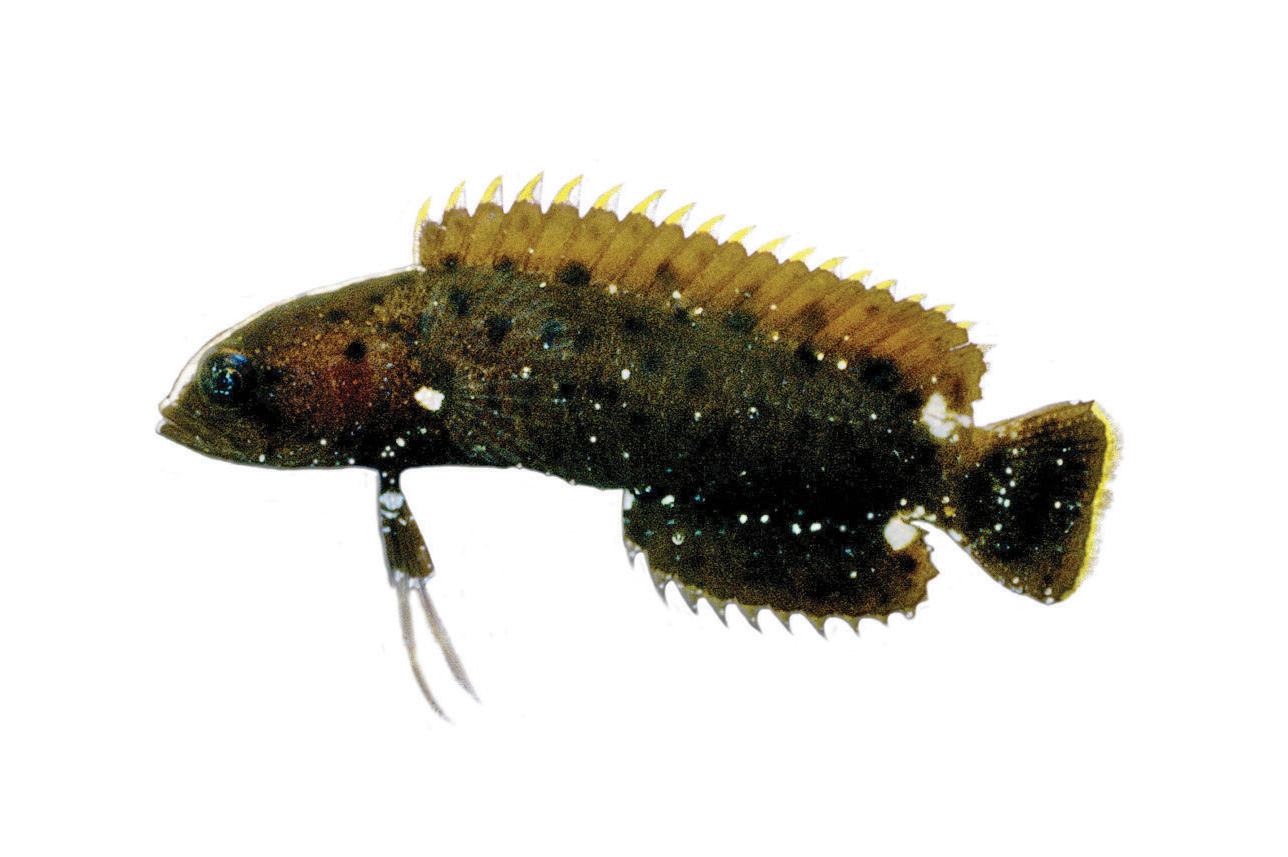
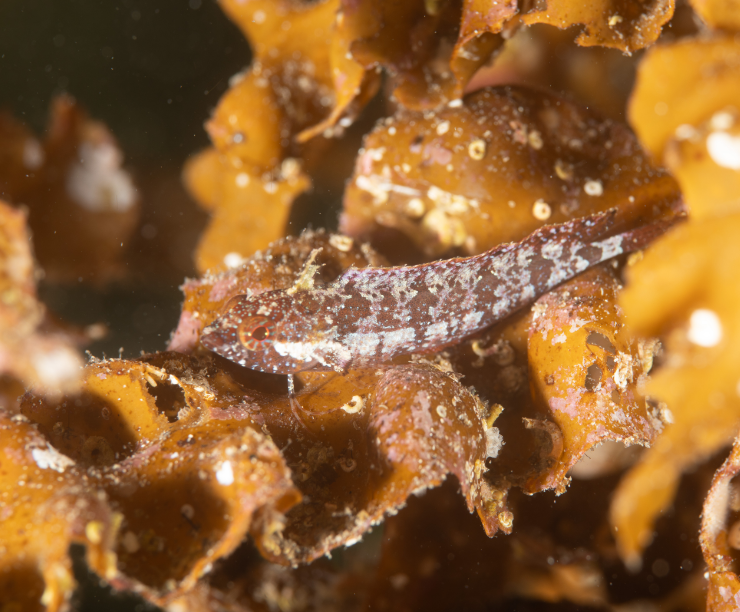
Species Exploration Tools: Interactive dashboard for exploring marine species through environmental DNA (eDNA) data with filtering by species name, IUCN Red List status, location, and date range
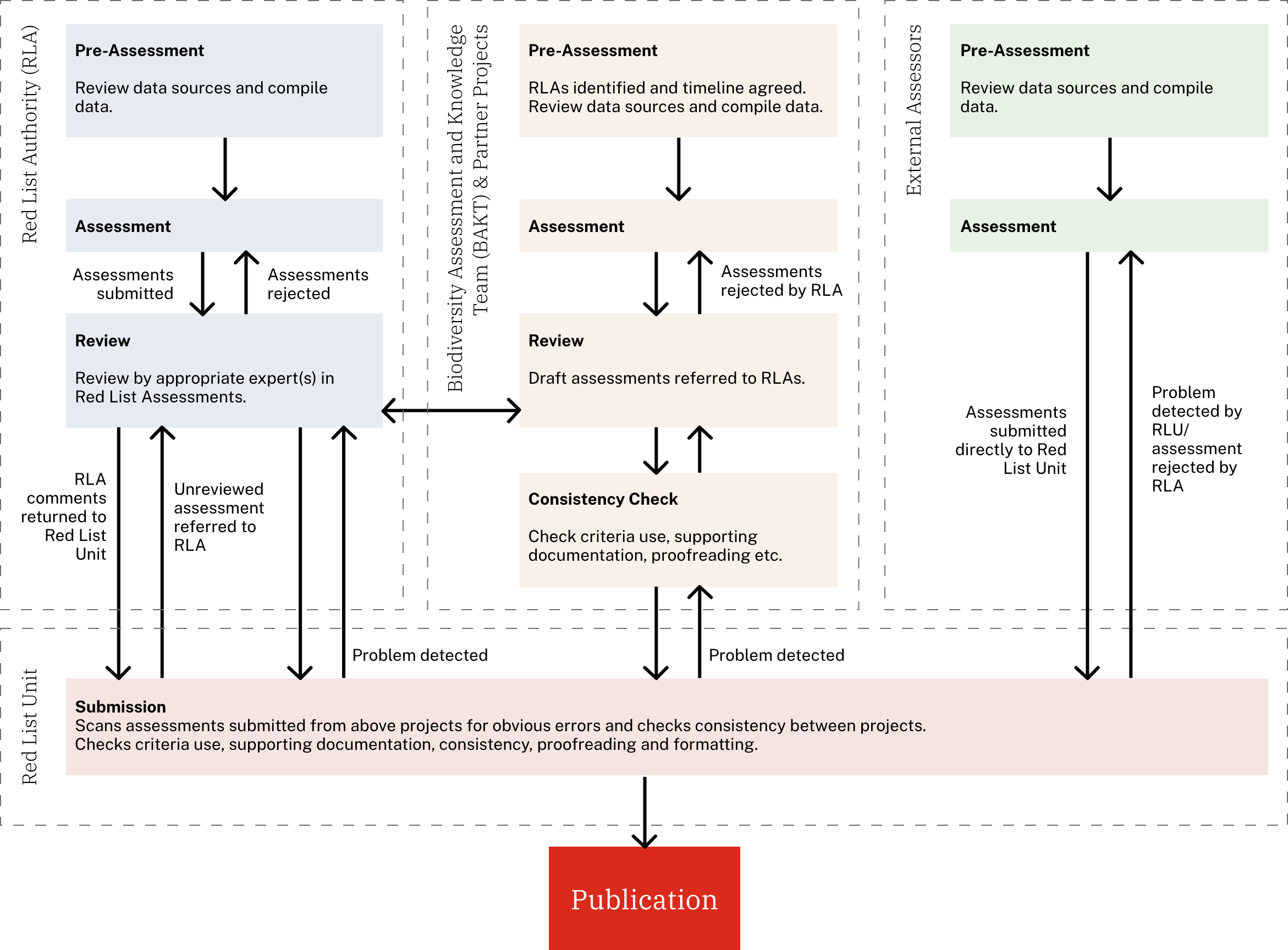
IUCN Red List Integration: Complete integration with IUCN Red List categories including Critically Endangered, Endangered, Vulnerable, Near Threatened, Least Concern, Data Deficient, and Not Evaluated species
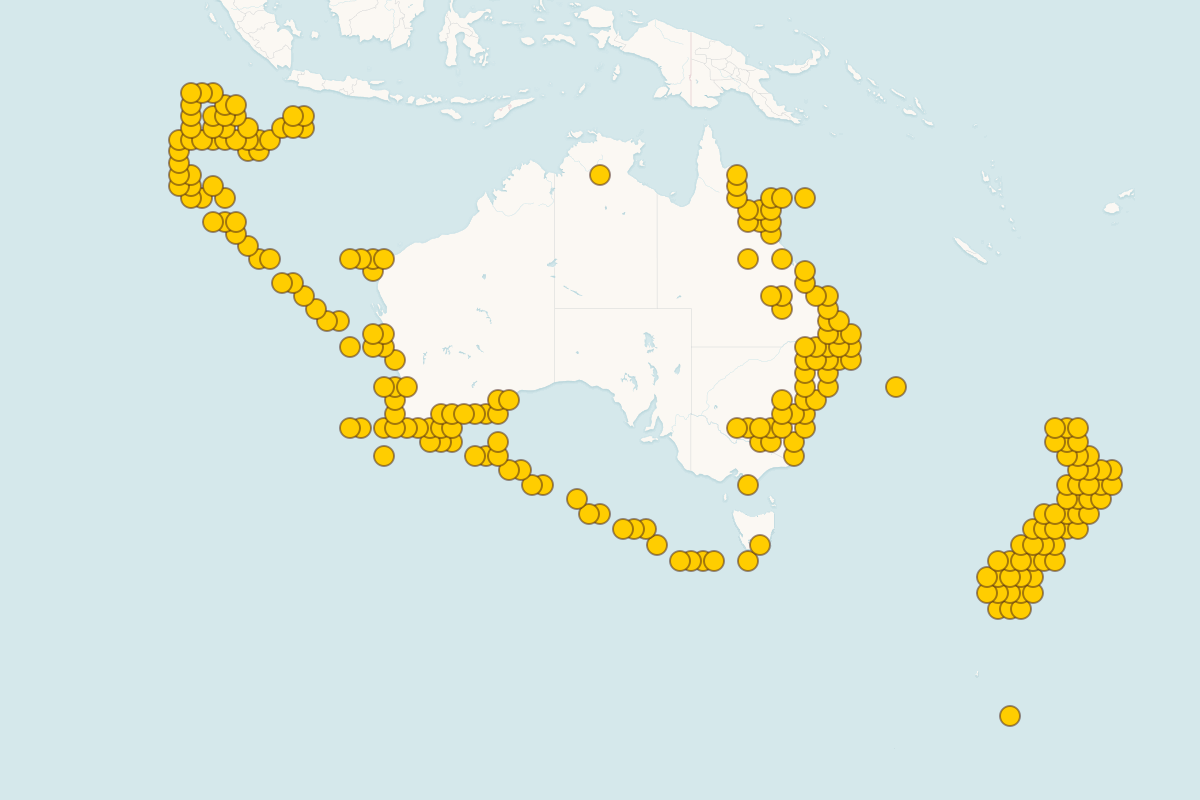
Interactive Mapping: Advanced MapLibre-based mapping system featuring marine park boundaries, species detection clusters, timeline animation controls, and customizable map views with satellite and street map options
Data Visualization: Dynamic charts including detections over time plots, species distribution maps, and comprehensive data tables showing eDNA sample locations and detection frequencies
Marine Parks Integration: Full integration with Parks Australia marine park network data, allowing exploration of species within protected areas and park-specific biodiversity insights
AI-Powered Data Chat: Revolutionary AI chat interface powered by Anthropic's Claude 4.5 Sonnet for natural language queries about the eDNA dataset, enabling real-time biodiversity insights
Assessment Tools: Specialized tools for IUCN assessors including data availability indicators, voucher sequence information, and base pair identity analysis for taxonomic verification
Comprehensive Dataset: Analysis of 6,000+ marine eDNA samples collected around Australia in collaboration with Parks Australia, covering diverse marine ecosystems
User Experience: Guided tours and interactive tutorials, responsive design with Bootstrap 5, real-time data filtering, and export capabilities for research and assessment workflows
Technical Infrastructure: Built on R Shiny with modern web technologies, Docker containerization, robust data processing pipeline using the targets framework, and SCSS-based styling system
This pilot dashboard represents the first-of-its-kind integration of environmental DNA data with IUCN Red List assessments, supporting efforts to rapidly incorporate eDNA evidence into global species extinction risk evaluations.